Chemical reaction networks drive many of the processes in living systems. Systems chemists aim to mimic this behavior by designing functional molecular systems. One major challenge in this context is to combine individual reactions into networks. Enzymes could play a key role in this, but there is currently no general and scalable route to construct enzymatic reaction networks
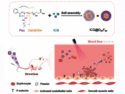
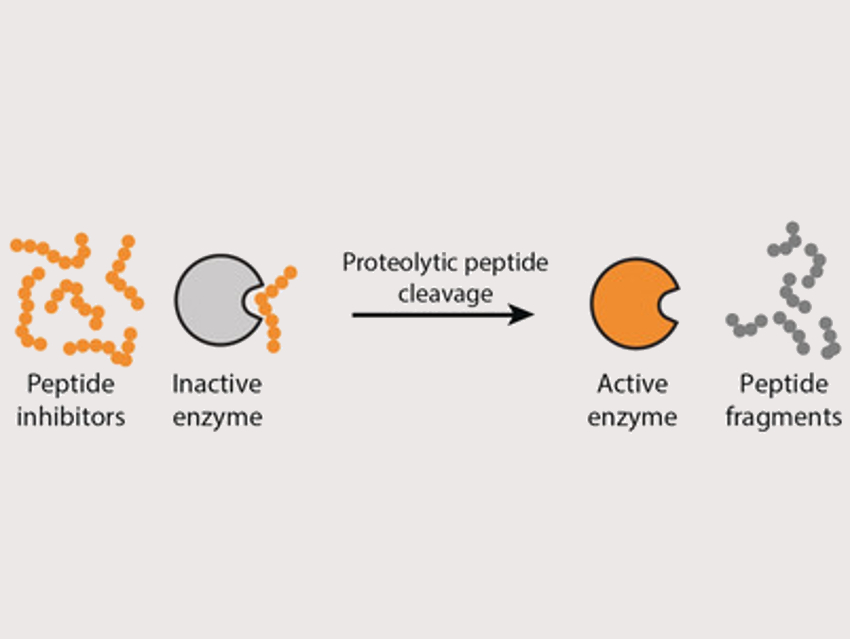
Wilhelm T. S. Huck, Radboud University, Nijmegen, The Netherlands, and colleagues have developed a way to control the activation of enzymes in simple networks. Regulating the activity of an enzyme is a crucial control element in such networks. The team used proteases (trypsin, chymotrypsin, or elastase) together with peptides that act as enzyme inhibitors and can be cleaved by these proteases. In the presence of the peptide inhibitor, most of the enzyme is in the inactive state (pictured left). When the inhibitor is cleaved by the proteases, the equilibrium shifts to the active enzyme (pictured right). This approach leads to auto‐activation of the different proteases.
The team also combined multiple enzymes in one experiment to study the properties of small reaction networks. In this setup, one enzyme can, for example, cleave the inhibitor of another, which leads to a coupling of the reactions. The researchers also developed an enzymatic off-switch, which turns precursors into active inhibitors to rapidly turn off enzymatic activity.
These findings allow the construction of larger reaction networks in a systematic way. The team’s ultimate goal is to build complex enzymatic reaction networks that show similar properties to living systems, e.g., sensing and signaling or information processing.
- Modular Design of Small Enzymatic Reaction Networks Based on Reversible and Cleavable Inhibitors,
Aleksandr A. Pogodaev, Cristina Lía Fernández Regueiro, Miglė Jakštaitė, Marijn J. Hollander, Wilhelm T. S. Huck,
Angew. Chem. Int. Ed. 2019.
https://doi.org/10.1002/anie.201907995