A variety of metabolic and neurological disorders could be more effectively treated with drugs that target class B G-protein-coupled receptors (GPCRs), as the only class B GPCR ligands currently available are molecules with poor pharmacokinetic properties. The design of small-molecule drugs specific for class B GPCRs would help solve this problem, but efforts in this vein have been hampered by a lack of structural information for class B receptors.
Crystallographic data are available for a number of class A GPCRs, but these differ considerably from the class B receptors, and thus modeling the 3D structure of a class B binding site with a class A template is a challenge. Didier Rognan, CNRS–University of Strasbourg, France, and colleagues performed the first successful structure-based identification of class B GPCR modulators.
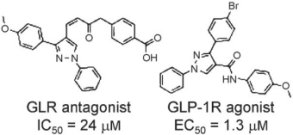
Starting from a 3D model of the glucagon receptor (GLR), a database of 1.9 million drug-like compounds was screened for similarity to existing non-competitive GLR antagonists and docked into the transmembrane cavity of GLR. The team’s new protein–ligand interaction fingerprint scoring method was used to filter and rank the docking poses. Among the hits, two compounds were validated as novel non-competitive GLR antagonists (pictured).
This customized screening approach opens new opportunities for designing class B GPCR ligands.
Images: © Wiley-VCH
- Structure-Based Discovery of Allosteric Modulators of Two Related Class B G-Protein-Coupled Receptors
C. de Graaf, C. Rein, D. Piwnica, F. Giordanetto, D. Rognan,
ChemMedChem 2011, 6(12).
DOI: 10.1002/cmdc.201100317