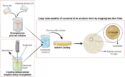
The CRISPR/Cas9 system is a useful tool for genome editing. It consists of single guide RNAs (sgRNA), which target specific sites in a DNA strand, and a Cas9 enzyme, which cuts the DNA at these sites. Some applications require the precise control of CRISPR/Cas9 in time and/or space. This can be achieved by regulating either sgRNA expression or Cas9 activity. However, existing methods suffer from limitations such as off-target effects or reduced editing capacities.
Ulrich Elling, Institute of Molecular Biotechnology of the Austrian Academy of Science (IMBA), Vienna, Austria, and colleagues have developed a method that can precisely control sgRNA expression. The team calls the approach CRISPR-Switch, short for “SgRNA With Induction/Termination by Cre Homologous recombination”. Cre is a recombinase enzyme that catalyzes a genetic recombination between two DNA recognition sites, called loxP sites. The team created a modified, loxP-containing sgRNA. This guide RNA can be controlled by regulating the Cre activity.
When the system is switched on, it allows a controlled, rapid induction of sgRNA activity. It can also be switched off again, which limits off-target effects. The team used the method to “knock out” different genes in a defined, programmed order. This allowed them to study the order of mutagenic events that lead to the development of glioblastoma (a type of brain cancer) in mice. According to the researchers, CRISPR-Switch substantially increases the versatility of gene editing.
- CRISPR-Switch regulates sgRNA activity by Cre recombination for sequential editing of two loci,
Krzysztof Chylinski, Maria Hubmann, Ruth E. Hanna, Connor Yanchus, Georg Michlits, Esther C. H. Uijttewaal, John Doench, Daniel Schramek, Ulrich Elling,
Nat. Commun. 2019.
https://doi.org/10.1038/s41467-019-13403-y