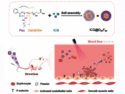
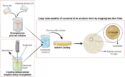
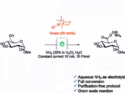
The crystal structure of M-PMV retroviral protease, which is linked to an AIDS-like virus, has been studied for over a decade, yet no definitive structure has been determined. Following the failure of a range of attempts to solve the structure by molecular replacement, David Baker and colleagues, University of Washington, USA, challenged players of the protein folding game Foldit to produce models of the protein.
The game’s players generated an accurate 3D model of the enzyme in just ten days. Baker’s team was then able to refine and determine the enzyme’s structure and identify surfaces on the molecule that are likely targets for deactivation. This technique combines the strengths of computers and humans to solve problems and provides new insights for the design of antiretroviral drugs.
- Crystal structure of a monomeric retroviral protease solved by protein folding game players
F. Khatib, F. DiMaio, S. Cooper, M. Kazmierczyk, M. Gilski, S. Krzywda, H. Zabranska, I. Pichova, J. Thompson, Z. Popović, M. Jaskolski, D. Baker,
Nat. Struct. Mol. Biol. 2011.
DOI: 10.1038/nsmb.2119 - Foldit Website