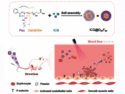
Biochemical events induce proteins to physically associate one to another, creating so-called protein-protein interactions. Targeting these interactions is becoming an attractive approach in drug discovery. This strategy, however, is very challenging because, as opposed to catalytic sites, interactions sites have unknown structures and different physico-chemical properties. Moreover, whereas synthetic macrocycles (MCs) have emerged as good candidates to target protein-protein interfaces, it still remains unclear how to design MCs for this purpose.
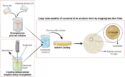
To overcome these limitations, Adrian Whitty, Boston University, USA, and co-workers examined a representative set of MC–protein complexes for which the structure is available. The scientists studied how MCs bind to their target proteins by analyzing the physicochemical features of the MCs regions directly involved in the binding. In addition, the scientists defined which MCs regions and atoms contribute to the binding and analyzed the geometry of the MCs ring relative to their target proteins. Altogether, this analysis allowed the researchers to develop specific guidelines to design large libraries of synthetic MCs endowed of good availability as well as strong binding to protein-protein interactions sites.
- Pharmacological chaperones stabilize retromer to limit APP processing,
Vincent J. Mecozzi, Diego E Berman, Sabrina Simoes, Chris Vetanovetz, Mehraj R. Awal, Vivek M. Patel, Remy T. Schneider, Gregory A. Petsko, Dagmar Ringe, Scott A Small,
Nature Chem. Biol. 2014, 10, 443–449.
DOI: 10.1038/nchembio.1508