RNA molecules perform many crucial functions in nearly every cellular pathway. Critical to controlling RNA function is its ability to fold into complex three-dimensional structures. Effort has been focused on understanding the structural components of RNA. Traditionally this has been done by one-RNA-at-a-time experiments. However, these conventional methods have now been married with genome-wide technologies to provide a systems view of RNA structure.
Miles Kubota, Dalen Chan, and Robert C. Spitale, University of California, Irvine, CA, USA, describe the use of chemical methods to probe RNA structure and how these methods have been merged with deep sequencing. They discuss how a combination of these two methods has revealed new understanding of RNA structure and function, particularly within a living cell, and what future challenges for transcriptome-wide RNA structure probing are.
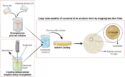
Various interatomic bindings determine how an RNA molecule folds. In the selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE) methode, a chemical reagent (the SHAPE probe) is mixed into a solution containing many thousands of RNA molecules. The SHAPE probe can chemically attack and modify every base of every RNA in the solution; the position of a modified base is then detected as a stop in an optimized primer extension reaction, followed by electrophoretic fragment separation. The speed with which the probe modifies the individual bases is an indication of how flexible the backbone of the RNA is at each base position. This flexibility is highly correlated with (but not wholly determined by) whether or not the base is paired to one of its complements. In SHAPE-Seq, SHAPE is extended by bar-code based multiplexing combined with RNA sequencing to be used in a high-throughput fashion.
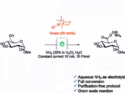
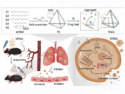
In hydroxyl radical cleavage, hydroxyl radicals damage the RNA depending on the local structure. Coupled with the high-throughput analytical capabilities of capillary electrophoresis this allows to quickly evaluate the structural variation of longer RNA sequences than was previously possible using standard gel electrophoresis. Additionally, the Multiplexed ·OH Cleavage Analysis, which uses anchored iron-radical cleavage for 3-D structure probing and modeling, has been married with deep sequencing. This makes it possible to measure RNA structure, transcriptome-wide, in three dimensions.
- RNA structure: Merging chemistry and genomics for a holistic perspective,
Miles Kubota, Dalen Chan, Robert C. Spitale,
BioEssays 2015.
DOI: 10.1002/bies.201300146