The chemical functionalization of proteins remains a challenge for biochemistry. Current technologies introduce labels at less abundant residues such as cysteine, tyrosine, or tryptophan. However, these residues could be inaccessible or their modification could lead to changes in the activity of the protein. Other residues like lysine (K), aspartate, or glutamate have also been used. Because of their abundance and relative accessibility, non-specific labeling at one or several sites usually occur.
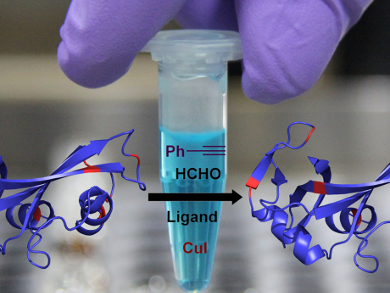
Vishal Rai, Indian Institute of Science Education and Research, Bhopal, India, and colleagues have devised a way to selectively label one lysine residue in a protein using a multicomponent mixture of formaldehyde, copper(I) iodide as a catalyst, 1,10-phenantroline as the ligand, and phenylacetylene as a label. The team optimized the reaction conditions and tested different catalysts and ligands using the protein RNAase A (13.7 kDa). They observed 90 % conversion of residue K31 against ten competing K residues under the optimitzed conditions.
To show the wide applicability of their method, the researchers singly labeled several proteins with varying sizes and numbers of K residues: Ubiquitin (8.5 kDa, 48 % conversion of K48, against 7 Ks), cytochrome c (12 kDa, 40 % conversion of K5, against 19 Ks), lactalbumin (14 kDa, 48 % conversion of K93, against 12 Ks), myoglobin (17 kDa, 45 % conversion of K147, against 19 Ks), and Subtilisin A (27 kDa, K264, 99 % conversion of K264 against 9 Ks). The team noted that the labeled lysine residues were all solvent-accessible based on the proteins’ 3D structures.
- Site-Selective Labeling of Native Proteins by a Multicomponent Approach,
Maheshwerreddy Chilamari, Landa Purushottam, Vishal Rai,
Chem. Eur. J. 2017.
DOI: 10.1002/chem.201605938