Natural products that contain nitrogen–nitrogen bonds display remarkable structural diversity and have a wide variety of biological activities. Several biochemical routes leading to enzymatic N–N bond formation have been investigated. However, the biosynthetic origin of naturally occurring pyrazole heterocycles has remained largely unexplored.
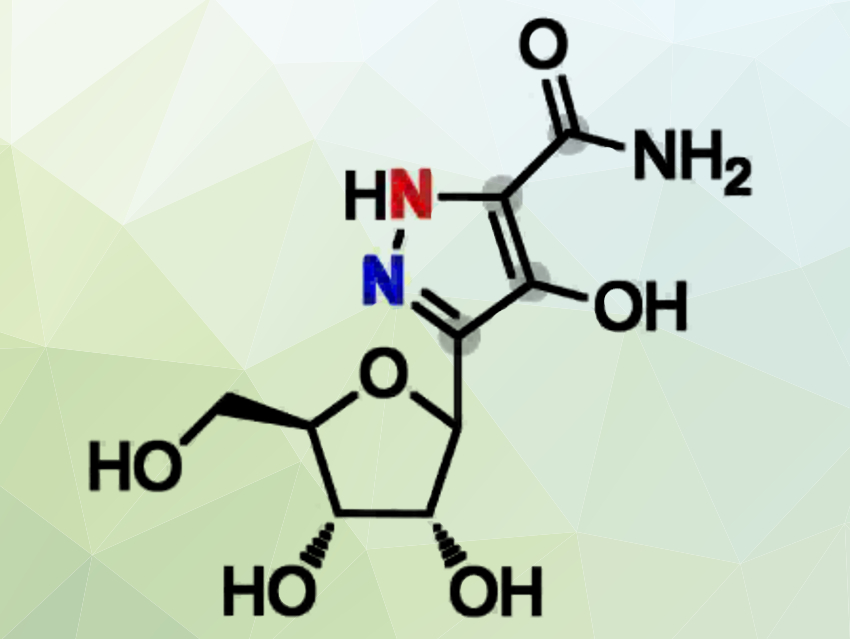
Yi-Ling Du, Zhejiang University, Hangzhou, China, and colleagues have pinpointed the biosynthetic gene cluster for pyrazomycin in Streptomyces candidus NRRL 3601. Pyrazomycin (pictured), also known as pyrazofurin, is an unusual pyrazole-containing C-nucleoside antibiotic with broad-spectrum antiviral and antitumor activities.
The team used isotope-labeled precursors to investigate the biosynthetic origin of the pyrazole moiety. Cells were fed with 13C/15N-labeled or unlabeled amino acid precursors and the isotopes were tracked in the products to find the origin of different parts of the target molecule. In addition, E. coli bacteria were genetically modified using the genes thought to be involved in the synthesis of pyrazomycin to confirm the role of the gene cluster—in particular, the gene pyrN, which encodes the enzyme PyrN.
The team found that the adjacent nitrogens in pyrazole ring derive from the ε-NH2 nitrogen of lysine and the α-NH2 nitrogen of glutamate, respectively. The enzyme which catalyzes this reaction, PyrN, is a new nitrogen–nitrogen bondforming enzyme. According to the researchers, this study helps to understand the early steps in pyrazomycin biosynthesis.
- The biosynthetic gene cluster of the C-nucleoside antibiotic pyrazomycin with a rare pyrazole moiety,
Guiyun Zhao, Shunyu Yao, Kristina W Rothchild, Tengfei Liu, Yu Liu, Jiazhang Lian, Hai-Yan He, Katherine S Ryan, Yi-Ling Du,
ChemBioChem 2019.
https://doi.org/10.1002/cbic.201900449