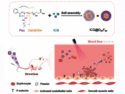
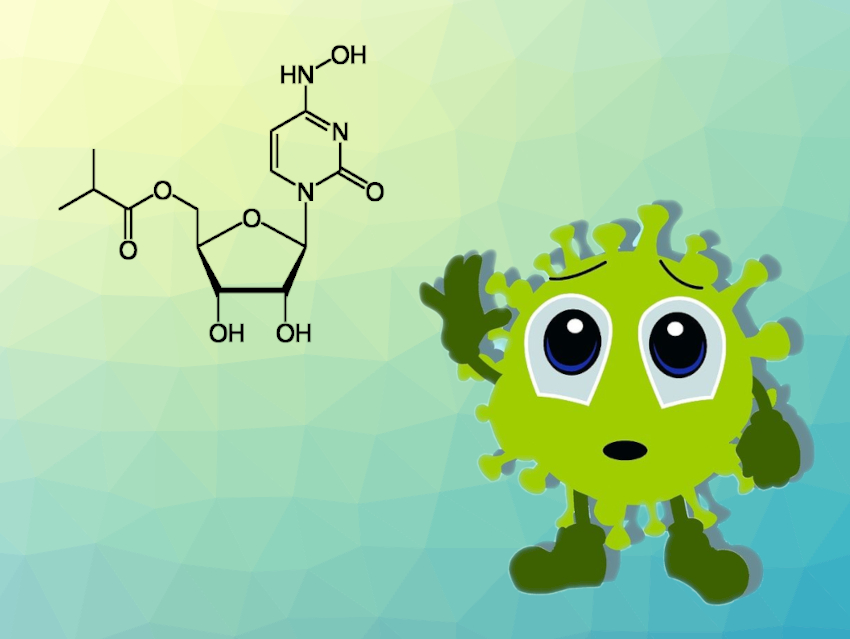
Nucleoside analogs are extremely promising molecules in the development of antiviral drugs. They act by interfering with viral RNA replication, and thus, hindering viral multiplication. Molnupiravir, an orally administered compound, is one such candidate and is currently in phase III clinical trials for use against SARS-CoV-2.
Claudia Höbartner, University of Würzburg, Germany, Patrick Cramer, Max Planck Institute for Biophysical Chemistry, Göttingen, Germany, and colleagues have established the molecular basis for its proposed mechanism of action.
Nucleoside Analogs
The concept behind nucleoside analogs as antiviral drugs is based on RNA polymerase incorrectly inserting them into a growing RNA strand as the virus attempts to replicate itself. Once supplied with faulty RNA, the virus might suffer from impaired protein synthesis or stalled RNA synthesis. This means it will either not work properly or not be able to replicate at all.
Although the capability of the virus to proofread RNA and correct errors has thwarted many of these attempts to harness nucleoside analogs, some positive results have been reported. For example, the drug remdesivir, also a nucleoside analog, received FDA approval for the treatment of COVID-19 in some patients. Remdesivir mimics adenosine triphosphate (ATP) after it has been converted to its active nucleotide form in the body.
After inserting this ATP analog into the growing RNA strand, viral RNA-dependent RNA polymerase stops further synthesis, because the cyano side group of the nucleoside analog blocks the next nucleoside insertion. However, despite this RNA-inhibiting activity, leading to decreasing viral load as shown in many in-vitro studies, clinical trials have revealed mixed results, first against Ebola, but also against SARS-CoV-2.
A New Hope
Molnupiravir is a new hope. Originally developed for use against influenza, promising results in human tissue, animals, and humans have elevated molnupiravir to the status of a highly interesting drug candidate in the fight against COVID-19.
Structurally, molnupiravir is the isopropyl ester of β-D–N4-hydroxycytidine. After intake, the body metabolizes it to form the active triphosphate form known as MTP. Research suggests that molnupiravir reduces the virus load because it is mutagenic. Treatment with the substance seems to introduce so many mutations that the virus can no longer replicate.
The team performed RNA synthesis and structural studies to reveal the origin of this mutagenesis. First, they performed templated RNA synthesis using RdRp, the RNA-dependent RNA polymerase belonging to SARS-CoV-2. They found that in contrast to remdesivir, MTP did not stall RNA synthesis. On the contrary, RdRp was unphased by the imposter nucleoside, continuing RNA synthesis for at least 11 nucleotides after MTP insertion. This means faulty RNA continues to be produced, rather than being corrected.
The Mutagenesis Hypothesis
Analyzing the new RNA sequences, the researchers discovered that MTP competed with both cytidine triphosphate and uridine triphosphate in the formation of the new RNA strand. The RNA polymerase placed the new nucleoside “M” opposite to guanosine, but also opposite to adenosine in the template strand. However, not all cytidines and uridines in the newly synthesized strand were replaced by M. The polymerase selected M for cytidine slightly less efficiently and inserted M instead of uridine at around half efficiency.
The team also used an M-containing RNA as a new template and discovered that RdRp incorporated either guanosine or adenosine opposite to M in a growing RNA strand. This random insertion of guanosine and adenosine implies that the original base pairing becomes increasingly disorganized with every new replication event. The researchers hypothesize that the growing mutation frequency causes so many errors in new RNA strands that the virus ultimately fails to replicate at all.
Base-Pairing Mimicry
Thermal melting experiments of RNA duplexes then revealed that M formed equally stable base pairs with both guanosine and adenosine. The researchers also looked at the structures of these base pairings. Using cryo-electron microscopy, they identified two hydrogen bridges connecting adenosine to the imino form of M. In contrast, guanosine formed three hydrogen bridges with the amino form of M.
This base-pair mimicry seems to work so well that M insertion manages to evade the viral defense system, and also explains the broad antiviral activity of molnupiravir. However, this effectiveness may cause problems for antiviral treatment in humans: scientists warn that molnupiravir could also defy human RNA polymerases. Regarding the mutagenicity of the compound, the researchers recommend more in-depth studies of the effects of molnupiravir on human cellular polymerases.
- Mechanism of molnupiravir-induced SARS-CoV-2 mutagenesis,
Florian Kabinger, Carina Stiller, Jana Schmitzová, Christian Dienemann, Goran Kokic, Hauke S. Hillen, Claudia Höbartner, Patrick Cramer,
Nat. Struct. Mol. Biol. 2021.
https://doi.org/10.1038/s41594-021-00651-0
Also of Interest
News: Coronavirus Drug Molnupiravir
- 05 October 2021
Molnupiravir significantly reduces risk of hospitalization and/or death, according to Merck
News: Improved Synthesis Route for Molnupiravir
- 04 August 2021
Efficient synthesis from cheap and easily available cytidine
Research Highlight: How Molnupiravir Causes SARS-CoV-2 to Mutate and Die
Mechanism might apply to various viral polymerases and could explain broad-spectrum antiviral activity of molnupiravir
News: High-Yielding Synthesis of Antiviral Drug Candidate
- 19 November 2020
Improved preparation of EIDD‐2801, currently in clinical trials for COVID-19 treatment
News: Oral Drug Blocks SARS-CoV-2 Transmission in Ferrets
- 09 December 2020
Repurposed ribonucleoside analogue inhibitor of influenza viruses