Random numbers can be produced by algorithms or generators based on unpredictable physical events. Robert N. Grass, Swiss Federal Institute of Technology (ETH) Zurich, Switzerland, and colleagues have developed the first true random number generator based on chemical synthesis. Their process consists of random DNA synthesis and sequencing: The randomly generated chain of nucleotides produces strings of bits, which can be processed and corrected for bias. According to the study, the speed and cost of this technique are comparable to current physical systems.
Why Random Numbers?
The world’s first random number generator was probably the dice. When throwing dice is insufficient for a particular randomizing task, modern people use algorithm-based random number generators. However, the random number generators that we are familiar with from app stores are not truly random. Their output depends on the input, i.e., the way the programmer has fed data into the system. Thus, algorithm-based generators are more accurately referred to as pseudo-random number generators.
In contrast, “true” random number generators use physical phenomena which are entirely independent of the operator. Radioactive decay is one such example. Another example is the atmospheric noise, which is used as the basis for many hardware-integrated true random number generators. These generators come into place when security is of utmost importance, for example, for secure data transmission and cryptography applications.
Might there also be a chemical way to generate true random numbers? For this, the researchers looked at DNA synthesis. Modern DNA synthesizers add the four DNA bases—adenine, thymine, cytosine, and guanine—to a growing DNA strand. The level of probability is roughly the same. This means that, after dozens of rounds, synthesis from a mixture of bases would rapidly produce an innumerable amount of random sequences.
Machine DNA Synthesis Has Its Biases
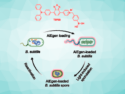
The team asked two commercial DNA synthesis labs to synthesize a library of random DNA sequences. The requested random strands were 64 nucleotides long, each strand being framed by defined primer and reverse primer sequences to enable polymerase chain reaction (PCR) replication.
The researchers then sequenced the mixtures obtained using commercial DNA sequencing machines. Their aim was to find out if the generated DNA library contained true random sequences or if there was any bias. Chemical synthesis has several sources of error. For example, all components must be present in exactly the same amounts in any given round. The team discovered that the distribution of bases along the strands was more unequal, or less random, when the four bases were not mixed before the synthesis step. However, when the bases were mixed before entering them in the synthesis, they were distributed more evenly along the strands.
The team also found two other biases. Firstly, guanine and thymine were more abundant in all sequences than adenine and cytosine. The researchers did not find a clear-cut explanation for this, but they observed that some bases showed preferences to attach to certain other ones during synthesis.
Secondly, the presence of guanine in the strands increased the longer the DNA strands grew, while that of thymine decreased. The researchers reasoned that guanine was prone to oxidation and the longer it rested in a strand, the more it tended to be replaced with thymine in a process called guanine–thymine transversion.
Correcting for Bias
To account for the biases observed, the team decided to apply algorithms that would strip bias from the output data rather than trying to change the synthesis conditions. First, the nucleotide sequences were converted to streams of bits, i.e., zero and one. Then, a rigorous algorithm processed the bit streams, discarding three-quarters of the raw data, but leaving bit streams that were perfectly random, as confirmed by randomness evaluation tests.
Speed and Storage
The researchers point out that it would not be speed, capacity, or costs that would hamper the development of true random number generation based on DNA synthesis. “DNA is manufactured by fully automated machines within 8.75 hours and can be obtained commercially for a price of approx. USD 100,” they state, calculating that the synthesized material obtained translated into a library of 1015 strands of individual DNA molecules, at an impressive speed of 225 gigabits of randomness per second.
Instead, the sticking point would be sequencing, which, even with the state-of-the-art technology available to the team, slowed down randomness generation to a moderate 300 kilobits per second. This rate placed the DNA-based random number generator in the middle of the pack when compared to currently used physical technologies.
However, DNA-based random number generation offers another advantage. The DNA molecule is extraordinarily stable and can store a large amount of information in a confined space. “DNA can be archived for millennia, preserving the generated randomness for generations to come,” the researchers conclude. That would make an interesting code to break for future archaeologists!
- DNA synthesis for true random number generation,
Linda C. Meiser, Julian Koch, Philipp L. Antkowiak, Wendelin J. Stark, Reinhard Heckel, Robert N. Grass,
Nat. Commun. 2020.
https://doi.org/10.1038/s41467-020-19757-y