Lipids are important molecules in living systems, where they serve as building blocks of cell membranes, participate in cell signaling, and store energy. The field of lipidomics, i.e., the study of pathways and networks of cellular lipids in biological systems, has revealed a vast diversity of lipid structures. Mass-spectrometry imaging can be used to investigate the localization of different lipids in biological systems. Nevertheless, the imaging of isomers that have the same molecular composition, but differ in the position of C=C bonds remains challenging.
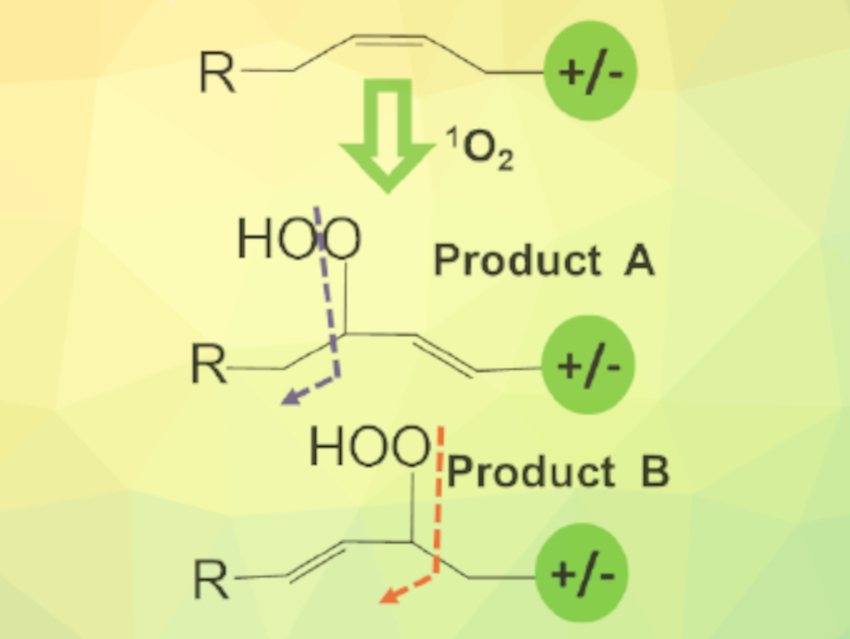
Julia Laskin, Purdue University, West Lafayette, IN, USA, and colleagues have developed a method for locating C=C bonds in lipids in imaging experiments. In their approach, lipids are extracted from the sample into a flowing solvent. The solvent containing the extracted lipids, as well as rose bengal as a photosensitizer, is then irradiated with a green laser pointer to generate singlet oxygen, which selectively reacts with C=C bonds in lipids (pictured). The hydroperoxide products of this reaction are broken up in a mass spectrometer, and the resulting fragments reveal the C=C location in the lipid.
This method is inexpensive due to the low-cost light source and photosensitizer, fast, and easy to implement on any mass spectrometer. It could improve the understanding of the role of lipid isomers in biological processes.
- Imaging and analysis of isomeric unsaturated lipids through online photochemical derivatization of C=C bonds,
Julia Laskin, Daisy Unsihuay, Pei Su, Hang Hu, Jiamin Qiu, Shihuan Kuang, Yingju Li, Xiaofei Sun, Sudhansu K. Dey,
Angew. Chem. Int. Ed. 2021.
https://doi.org/10.1002/anie.202016734