Glioblastoma is the most common and aggressive solid brain tumor with a mean life expectancy of only 12 to 15 months in patients. Standard medical care includes surgical resection, radiation, and systemic chemotherapy. Unfortunately, due to the diffuse growth and highly infiltrative capacities of the cancer cells, standard therapy is often not enough and patients suffer from tumor relapse.
To date, major treatment obstacles of a systemic chemotherapeutic approach are the natural diffusion barriers of solid tumors preventing the delivery of accurate doses of cancer drugs. In addition, currently used strategies such as pressure-gradient infusion of drugs are accompanied by serious side-effects that can cause long-lasting damage to the healthy brain.
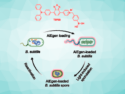
Ravi V. Bellamkonda, Duke University, Durham, NC, USA, and colleagues have designed a drug delivery strategy based on a non-pathogenic strain of the gut bacteria Salmonella typhimurium. The advantage of this approach is the higher motility that bacterial vehicles can achieve within the tumor. This increased motility leads to an improved delivery of the anticancer treatment. The scientists exploit a biological feature of tumor cells, e.g. their richness in purines, to target purine-deficient bacterial carriers more specifically to tumor cells.
To eliminate tumor cells, the bacteria are loaded with a specific killcode in form of DNA, coding for tumoricidal proteins such as the tumor suppressor protein p53 and/or Azurin. The expression of those proteins was put under the control of a promoter that will be turned on under hypoxic conditions, which reflects the microenvironment present within brain tumors. The researchers could demonstrate that tumor-bearing mice treated with bacterial vesicles showed an overall survival benefit of 19 % compared with non-treated mice.
- Bacterial Carriers for Glioblastoma Therapy,
Nalini Mehta, Johnathan G. Lyon, Ketki Patil, Nassir Mokarram, Christine Kim, Ravi V. Bellamkonda,
Mol. Ther. – Oncolytics 2017, 4, 1–17.
DOI: 10.1016/j.omto.2016.12.003