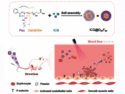
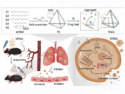
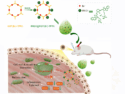
The coronavirus SARS-CoV-2 causes the current outbreak of the respiratory disease COVID-19. Researchers worldwide are focusing their efforts on investigating this virus and finding treatment options. For this research, it is important to understand how the proteins of the virus work and what drug candidates could bind to them. Computer simulations can help with this.
However, since proteins are large and complicated, with many degrees of freedom, their simulations can be very complex and need a large amount of computing time—or input from the puzzle-solving intuition of humans. You can help with both if you own a computer: by helping to solve protein-structure puzzles yourself or by donating some of your computing power.
Play a Game about Proteins
The scientific computer game Foldit uses the input of human players to figure out the final structure that a protein folds into. No scientific training or prior knowledge is necessary to play the game. Players are presented with the general structure of a protein and can then arrange it into a stable structure. They get points for properties that generally improve stability, such as a tight packing or avoiding interactions of hydrophobic sidechains with the water that usually surrounds a protein.
The project has released a coronavirus binder design challenge, where players can help to discover new antiviral drugs against SARS-CoV-2. In the puzzles of this challenge, players design and fold a new protein that can bind to the spike protein of SARS-CoV-2, which the virus uses to enter human cells. These antiviral proteins could then be useful to block infection. The most promising solutions will be tested in real life at the University of Washington’s Institute for Protein Design in Seattle, USA.
Donate Your Computing Power
If you do not want to fold proteins “by hand”, you can still help. Projects such as Folding@home use the distributed computing power of large numbers of computers all over the world to simulate, e.g., protein dynamics in collaboration with a number of research labs. Users can download software that allows their unused computing power to benefit the project, while they can keep using the computer normally. Together, the donated computing power rivals that of the largest supercomputers.
Folding@home currently runs simulations on the dynamics of SARS-CoV-2 proteins to find new therapeutic opportunities and sites that drugs could target. Users can contribute by downloading the Folding@home software.
Also of Interest
- Collection: SARS-CoV-2 Virus
What we know about the new coronavirus and COVID-19 - LitCovid
Curated literature hub for tracking up-to-date scientific information about COVID-19 - Many publishers and other entities have signed a joint statement to ensure that COVID-19 research findings and data are shared rapidly and openly