Since the SARS-CoV-2 virus, which causes COVID-19, was first detected, several new variants have emerged, including those first detected in the UK (B.1.1.7), South Africa (B.1.351), Brazil (P.1), and India (B.1.617). Some of these variants appear to be more transmissible, and thus, spread rapidly. In addition, there is the concern that the mutations present in these viruses could reduce the effectiveness of current vaccines, antibody therapies, or natural immunity.
The variant B.1.351 that first emerged in South Africa has three mutations. Two of them (N501Y and E484K) can enhance binding between the receptor binding domain (RBD) of the coronavirus spike protein and the ACE2 receptor on human cells, which helps the virus to enter host cells. However, the third mutation (K417N, a lysine to asparagine mutation at position 417) is less straightforward to explain because it removes a favorable interaction between the RBD and ACE2.
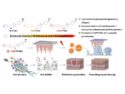
Binquan Luan and Tien Huynh, IBM Thomas J. Watson Research, Yorktown Heights, NY, USA, searched for potential benefits of the K417N mutation that could have caused the coronavirus to evolve along this path. The researchers used molecular dynamics (MD) simulations to analyze the effects of the K417N mutation in variant B.1.351. First, they modeled the binding between the original SARS-CoV-2 RBD and ACE2, as well as the binding between the RBD and CB6, a neutralizing antibody isolated from a recovered COVID-19 patient. They found that the original amino acid, a lysine, at position 417 in the RBD interacts more strongly with CB6 than with ACE2, which is consistent with the antibody’s therapeutic efficacy.
Then, the team modeled the binding properties of the K417N variant, in which the lysine is changed to an asparagine. This mutation reduces the strength of binding between the RBD and ACE2, but it also decreases the RBD’s binding to CB6 and several other human antibodies to a much greater extent. Thus, the variant B.1.351 seems to have sacrificed tight binding to ACE2 at this site for the ability to evade the immune system. According to the researchers, these results could be useful for enhancing the protection provided by COVID-19 vaccines and therapies.
- Insights into SARS-CoV-2’s Mutations for Evading Human Antibodies: Sacrifice and Survival,
Binquan Luan, Tien Huynh,
J. Med. Chem. 2021.
https://doi.org/10.1021/acs.jmedchem.1c00311
Also of Interest
- Collection: SARS-CoV-2 Virus
What we know about the new coronavirus and COVID-19