Chinese Hamster Ovary (CHO) cells are commonly used in bioproduction of high quality therapeutic proteins compatible with humans. Genome editing tools such as TALENs, ZFNs, and meganucleases used to modify CHO cells are expensive and time-consuming, creating demand for cheaper, faster, and efficient alternatives.
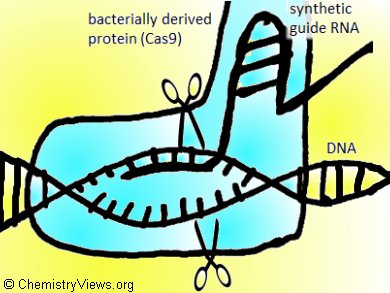
Michael Betenbaugh, John Hopkins University, Baltimore, MD, USA, Alex Toftgaard Nielsen, Technical University of Denmark, Lyngby, and their groups demonstrated for the first time the application and efficacy of a CHO codon-optimized CRISPR RNA-guided endonuclease Cas9 in CHO-K1 cells. They did this by successfully generating a large number of site-specific disruptions in COSMC and FUT8 glycosylation genes.
The researchers also developed the first web-based bioinformatics tool “CRISPy” for optimal single guide RNA (sgRNA) design in CHO cells. So far, there was no tool available to search for targets in CHO cells.
The demonstrated efficacy of the CRISPR Cas9 technology in the CHO cells combined with the new web tool offer an efficient, fast, and low cost alternative to above mentioned genetic manipulation tools. Both new approaches have the potential to accelerate metabolic engineering and synthetic biology efforts in the CHO cells.
- Accelerating genome editing in CHO cells using CRISPR Cas9 and CRISPy, a web-based target finding tool,
C. Ronda, L. E. Pedersen, H. G. Hansen, T. B. Kallehauge, M. J. Betenbaugh, A.T. Nielsen, H. F. Kildegaard,
Biotechnol. Bioeng. 2014.
DOI: 10.1002/bit.25233