Treatments against infections with SARS-CoV-2 and its variants of concern are an important research target. Drugs that target the important proteins in the viral life cycle can be useful for this. The main protease is one such important protein and acts as the target for drugs like paxlovid and molnupiravir. Using other viral proteins as drug targets could unlock even more effective ways to counter the virus.
With this goal in mind, Harald Schwalbe, Goethe University Frankfurt, Frankfurt am Main, Germany, and colleagues have developed protocols for the production and NMR characterization of more than 80 % of all SARS-CoV-2 proteins. The team then used NMR spectra to screen large sets of small organic molecules (a fragment library), testing their binding to the proteins and identifying lead compounds that can be further developed into drug-like molecules.
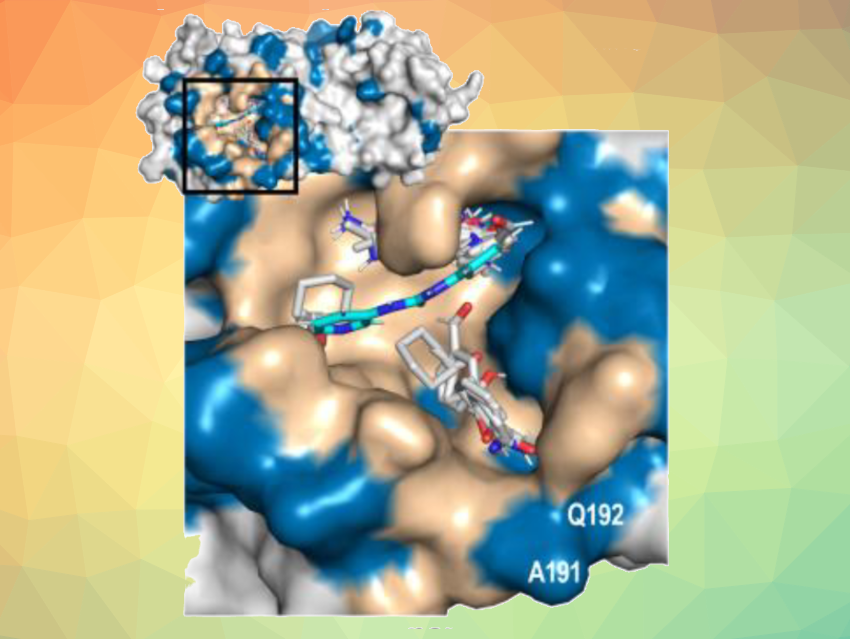
The team screened a fragment library composed of 768 highly diverse fragments against 25 SARS-CoV-2 proteins, providing information on 19,200 possible protein–ligand interactions. They identified 311 hits, i.e., fragments that bind to SARS-CoV-2 proteins. The researchers then developed a computational method to discover the binding pocket on the protein for these fragments (example pictured)
The obtained data could be highly valuable to support medicinal chemistry research aimed at developing new drugs against SARS-CoV-2 protein targets.
- Comprehensive Fragment Screening of the SARS‐CoV‐2 Proteome Explores Novel Chemical Space for Drug Development,
Hannes Berg et al.,
Angew. Chem. Int. Ed. 2022.
https://doi.org/10.1002/anie.202205858
Also of Interest
- Collection: SARS-CoV-2 Virus
What we know about the new coronavirus and COVID-19