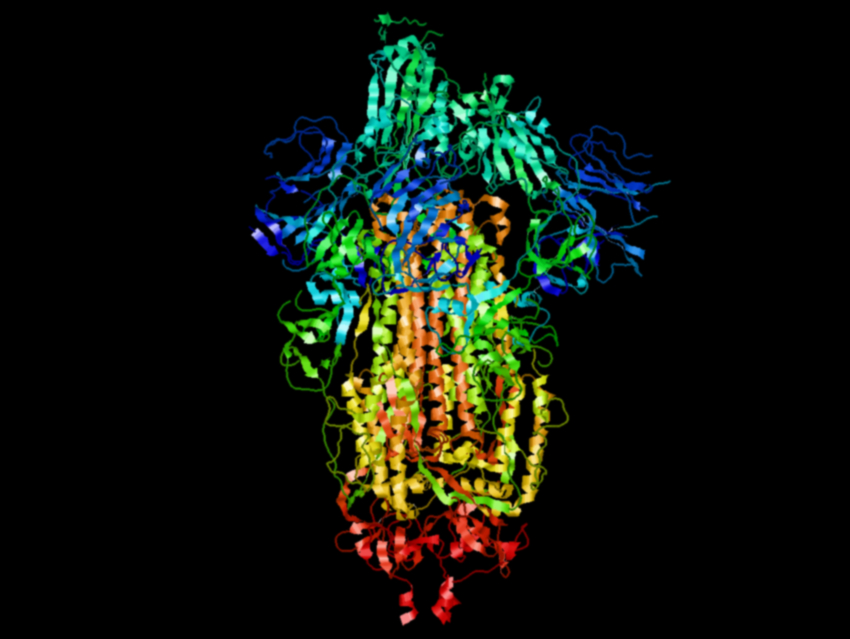
Coronaviruses, e.g., SARS-CoV-2 which causes the current COVID-19 pandemic, have glycoprotein “spikes” on their surface. These spikes help the virus to enter host cells by fusing the virus with cell membranes. The SARS-CoV-2 spike binds to a receptor on the surface of the host cells’ membrane—the angiotensin-converting enzyme 2 (ACE2). When it binds, the spike’s confirmation rearranges from a metastable prefusion structure to a stable postfusion structure.
The prefusion structure could be a target for the development of COVID-19 vaccines and treatments. However, this conformation of the protein is difficult to produce by recombinant expression, i.e., using cells that have been modified with foreign DNA. SARS-CoV-2 spike proteins that are stabilized in their prefusion configuration would, thus, be useful to improve protein yields.
Jennifer A. Maynard, Ilya J. Finkelstein, Jason S. McLellan, and colleagues, University of Texas, Austin, USA, have designed modified versions of the SARS-CoV-2 spike protein to find more stable variants that can be expressed at higher levels. The team analyzed the cryogenic electron microscopy (cryo-EM) structure of the spike protein and chose substitutions that should make the structure more stable, such as additional disulfide bonds, salt bridges, proline residues, and hydrophobic residues that can fill internal cavities. The team obtained 100 variants, each changed in a single position, and tested their expression in FreeStyle 293-F cells.
The researchers found that 26 of these single substitutions led to improved protein yields. Then they combined the best-performing substitutions to further optimize the stability and expression of the protein. They found that a variant with six proline substitutions, which they call HexaPro, is particularly promising. It has almost ten-fold higher expression than the parent spike protein, as well as increased thermostability. The researchers determined the cryo-EM structure of HexaPro (pictured) with a resolution of 3.2 Å and found that it retains the prefusion spike conformation.
- Structure-based design of prefusion-stabilized SARS-CoV-2 spikes,
Ching-Lin Hsieh, Jory A. Goldsmith, Jeffrey M. Schaub, Andrea M. DiVenere, Hung-Che Kuo, Kamyab Javanmardi, Kevin C. Le, Daniel Wrapp, Alison G. Lee, Yutong Liu, Chia-Wei Chou, Patrick O. Byrne, Christy K. Hjorth, Nicole V. Johnson, John Ludes-Meyers, Annalee W. Nguyen, Juyeon Park, Nianshuang Wang, Dzifa Amengor, Jason J. Lavinder, Gregory C. Ippolito, Jennifer A. Maynard, Ilya J. Finkelstein, Jason S. McLellan,
Science 2020.
https://doi.org/10.1126/science.abd0826
Also of Interest
- Collection: SARS-CoV-2 Virus
What we know about the new coronavirus and COVID-19 - LitCovid
Curated literature hub for tracking up-to-date scientific information about COVID-19 - Many publishers and other entities have signed a joint statement to ensure that COVID-19 research findings and data are shared rapidly and openly