Our genome is a chemical text written in our DNA using four letters. According to the blueprints it contains, thousands of different proteins are built from 20 amino acids in the cells of all living beings. The mechanism by which this four-letter DNA is transformed into a twenty-letter amino acid text is known as the genetic code.
Many brilliant minds worked on deciphering this code, and the goal was thought to have been achieved over sixty years ago: Crick, Griffith, and Orgel published a code so captivatingly beautiful that it simply must have been right [1-3].
Then came a surprise: Marshall W. Nirenberg and his German postdoctoral assistant J. Heinrich Matthaei successfully undertook the experimental confirmation of the “code without commas“. A relatively simple experiment proved that nature, unfortunately, does not use this brilliant code, preferring to take less elegant routes of its own.
Let’s take a closer look at this seeming chemical irrationality of nature.
6. UUU Codons
The Experiment
Marshall W. Nirenberg and his German postdoctoral student J. Heinrich Matthaei carried out their experiments in a cell-free system. They ground up harmless gut bacteria (Escherichia coli) with fine aluminum oxide. After filtering off the solid components, it was possible to maintain protein synthesis for hours after the addition of amino acids.
However, Nirenberg and Matthaei were not interested in protein synthesis; they had a completely different way of using this cell-free system. First, they destroyed the natural DNA that was present. DNA can be fully hydrolyzed—and thus, switched off—by addition of the enzyme deoxyribonuclease (DNAase). The result is that protein synthesis comes to a complete halt after a few minutes. Without DNA, no short-lived mRNA can be formed, and without mRNA, there is no synthetic template for making proteins.
Nirenberg and Matthaei had the ingenious idea of replacing the missing mRNA with a synthetic RNA to restart the protein synthesis. However, the only synthetically accessible RNA at the time was the very primitive poly(U) [13], which consists of only a single nucleotide, uridine diphosphate.
On Saturday, May 27, 1961, at 3:00 (in the morning!), Matthaei carried out his crucial experiment at the National Institutes of Health in Bethesda, USA [14]: He ground up E. coli bacteria, filtered off the solid components, and added DNAase to the filtrate to destroy the natural DNA. He then added synthetic poly(U) (an RNA with a base sequence of …UUUUUUU…) and phenylalanine that had been labeled with the radioactive isotope 14C. Protein synthesis started up immediately and, within an hour, the resulting protein was precipitated out and its radioactivity measured.
The result was impressive: Whereas control experiments carried out in parallel without poly(U) resulted in 40 to 60 radioactive decays per minute and milligram of protein, the use of poly(U) produced an exorbitant 39,800 decays. The result was unambiguous: the protein synthesis mechanism does not differentiate between synthetic RNA and natural mRNA. Any RNA sequence is blindly transcribed into a protein.
Poly(U) contains only UUU codons, which could now be unequivocally associated with phenylalanine (Phe). UUU was, thus, the first building block of the genetic code to be deciphered. This was a sensation, not only because a codon had been decoded, but because it was one with three identical bases. Precisely these codons—AAA, CCC, GGG, and UUU—are meaningless in the code without commas and are not supposed to code for any amino acid (see Chapter 4 in Part 1). Nirenberg and Matthaeis’ experimental result, thus, meant the end for the code without commas.
The Bomb Goes Off
No one noticed this at first, because Nirenberg was not in the exclusive RNA Tie Club and otherwise not part of the scientific establishment. The bomb only went off at the International Congress for Biochemistry in August 1961, in Moscow, Russia, with all the notables in the field attending. Nirenberg’s fifteen-minute presentation about “The Dependence of Cell-Free Protein Synthesis in E. Coli upon Naturally Occurring or Synthetic Polyribonucleotides” raised little interest and took place in a small room with sparse attendance.
Renowned biochemist Matthew Meselson reported: “I think that no one whom I knew well heard it. I heard the talk and was bowled over by it. It is dreadful snobbery that a presenter either belongs to important circles and is known, or one doesn’t know him, and it is unlikely that his work is important. And here was some guy named Marshall Nirenberg; his results were unlikely to be correct, because he wasn’t in the club. And nobody bothered to be there to hear him. Anyway, I was bowled over by the results.”
Meselson subsequently spoke to Crick about the sensational results, after which Nierenberg was invited to repeat his presentation in a larger, fully occupied room. Meselson also reported on this event: “After the splendid presentation, I went to Nirenberg, embraced him, and congratulated him. This is something I never do; it is not in my nature. However, I had the feeling that he had been treated unfairly and had a bit of a bad conscience.”
Because it was now known that phenylalanine was unequivocally assigned to the UUU codon, the code without commas had to be abandoned. Every amino acid must, thus, be encoded by one or several of the maximum of 64 codons. Optimistically, Crick and Brenner predicted that it would only take one year to fully solve the genetic code [15]. They were very mistaken: it took over five years.
The Chemistry of RNA Is Difficult
Why did it take so long? The answer is simple: The chemistry of RNA is difficult. To unequivocally assign a codon to an amino acid, the three corresponding nucleotides need to be attached to each other. The synthesis of poly(U) and the others was relatively easy because one “only” needed to hook uridine phosphates together using polynucleotide phosphorylase.
However, if given a mixture of different nucleotides, the enzyme hooks them up randomly, with no defined sequence. This limitation only allowed Nirenberg and Ochoa, working in parallel between 1960 and 1962, to determine the overall composition of the codons, but not the internal sequence of the three nucleotides in each (see Tab. 3).
|
Table 3. Determining the overall composition of codons. |
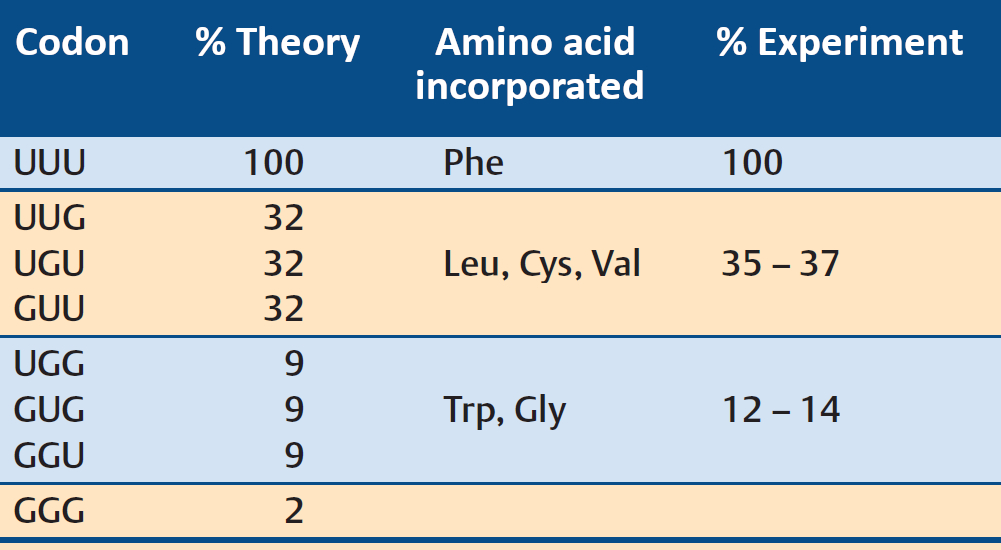 |
Determination of the overall composition of a codon can be illustrated with a typical Nirenberg experiment [16]: In an RNA made from 75 % U and 25 % G, one obtains the distribution shown in Tab. 3 for the relative frequency of the possible codons. Comparison of the calculated base distribution in the codons and the relative quantities of integrated amino acids allows amino acids to be assigned to specific nucleotide compositions of some codons, but not the sequence of their bases. In this case, they only determined that leucine, cysteine, and valine are all coded by codons with a composition of (U2G), but the internal sequence was undetermined.
7. RNAs with Defined Nucleotide Sequences
Nirenberg and Leder
Further elucidation of the genetic code required RNAs with defined nucleotide sequences. Two research groups solved this problem in different ways. Nirenberg and his postdoctoral assistant Philip Leder [16] started with the observation that both mRNA and the amino-acid-laden tRNA must be bound to the ribosomes during protein synthesis.
They developed the following experimental protocol: They first added the 20 amino acids, one of which was labeled with 14C, to a cell-free system. Then, in place of a polymeric RNA, they added an easily synthesized codon, just three nucleotides attached to each other in a known sequence. This codon bound to the ribosomes and, at the same time, to the amino-acid-laden tRNA corresponding to the codon. The ribosomes were filtered off, and their radioactivity was measured. If the radioactivity was high, then the labeled amino acid corresponded to the codon being tested. This was a truly elegant direct decoding method.
Gobind Khorana
At the same time, Indian-American chemist Gobind Khorana developed an alternative decoding technique. He was a leading expert in the organic chemical synthesis of short DNA and RNA segments and was convinced that “the responsibility for the complete decoding of the genetic code was in the hands of chemists.” Indeed, he successfully developed a synthetic method for making RNA with repeating sequences.
By synthesizing different RNAs with constantly repeating building blocks, Khorana was able to decipher a large portion of the genetic code. We can appreciate his impressive reasoning by looking at two of his experiments.
Khorana synthesized an RNA with constantly repeating AC building blocks, poly(AC). Independent of the starting point, the RNA (…ACACACACACACACACAC…) can only be read as (…ACA-CAC-ACA-CAC-ACA-…). The protein synthesized by this RNA must have two alternating amino acids. Experimentally, Khorana found the following amino acid sequence: (…Thr-His-Thr-His- Thr-His-Thr-His- Thr-His-…). The conclusion from this is that either ACA encodes threonine and CAC histidine, or vice versa.
Khorana then synthesized poly(AAC). This RNA, (…ACAACAACAACAACAACA…), can be read in codons three different ways:
- …ACA ACA ACA ACA ACA ACA ACA ACA ACA…
- …A CAA CAA CAA CAA CAA CAA CAA CAA CA…
- …AC AAC AAC AAC AAC AAC AAC AAC AAC A…
In the experiment, poly(threonine), poly(asparagine), and poly(glutamine) were detected. Combining the results of both experiments leads to the conclusion that ACA encodes threonine, and thus, CAC encodes histidine.
Nirenberg and Khorana’s research groups complemented and verified each other’s work and by combining all the data and clarifying ambiguous results, they were able to fully decipher the genetic code by 1966 (see Tab. 1). Both scientists were awarded the Nobel Prize in Physiology or Medicine in 1968. The third researcher awarded that prize in 1968 was Robert Holley, who co-discovered tRNA and was the first to fully determine its chemical structure.
List of Scientists
.jpg) |
Marshall W. Nirenberg (1927, New York, USA – 2010, New York, USA) |
 |
J. Heinrich Matthaei (*1929, Bonn, Germany) |
 |
Matthew Stanley Meselson (*1930, Denver, Colorado, USA) (Photo: modified from Janet Montgomery, wikimedia commons, CC BY-SA 4.0) |
 |
Francis Harry Compton Crick (1916, Northamptonshire, UK – 2004, San Diego, CA, USA) (Photo: Marc Lieberman, wikimedia commons, CC BY 2.5) |
 |
Severo Ochoa de Albornoz (1905, Luarca, Spain –1993, Madrid, Spain) Physician and biochemist, Nobel Prize for Physiology or Medicine in 1959 with Arthur Kornberg “for their discovery of the mechanisms in the biological synthesis of ribonucleic acid and deoxyribonucleic acid” |
 |
Philip Leder (1934, Washington, D.C, USA – 2020, Chestnut Hill, MA, USA) Geneticist, known, e.g., for his work with Marshall Nirenberg in the elucidation of the genetic code |
 |
Har Gobind Khorana (1922, Raipur, Punjab – 2011, Concord, MA, USA) Biochemist, 1968 Nobel Prize in Physiology or Medicine with Marshall W. Nirenberg and Robert W. Holley “for their interpretation of the genetic code and its function in protein synthesis”
|
 |
Robert William Holley (1922, Urbana, IL, USA – 1993, Los Gatos, CA, USA) Biochemist, 1968 Nobel Prize in Physiology or Medicine with Har Gobind Khorana and Marshall W. Nirenberg “for their interpretation of the genetic code and its function in protein synthesis”
|
References
[13] S. Ochoa, Enzymatic synthesis of ribonucleic acid, Nobel Lecture, December 11, 1959.
[14] M. Nirenberg, J. H. Matthaei, The Dependence of Cell-free Protein Synthesis in E. Coli upon Naturally Occurring or Synthetic Polyribonucleotides, Proc. Nat. Acad. Sci. USA 1961, 47, 1588–1602. https://doi.org/10.1073/pnas.47.10.1588
[15] F. H. C. Crick at al., General Nature of the Genetic Code for Proteins, Nature 1961, 192, 1227–1232. https://doi.org/10.1038/1921227a0
[16] P. Leder, M. Nirenberg, RNA Codewords And Protein Synthesis. II. Nucleotide Sequence Of A Valine RNA Codeword, Proc. Nat. Acad. Sci. USA 1964, 52, 420–427. https://doi.org/10.1073/pnas.52.2.420
The article has been published in German as:
- Die schönste falsche Theorie der Biochemie. Aus 4 mach’ 20, oder 21, oder 22, oder ….,
Klaus Roth,
Chem. unserer Zeit 2007, 41, 448–458.
https://doi.org/10.1002/ciuz.200700446
and was translated by Caroll Pohl-Ferry.
Deciphering the Genetic Code: The Most Beautiful False Theory in Biochemistry – Part 1
Countless scientists contributed to its clarification with brilliance, ingenuity, intuition, and luck
Deciphering the Genetic Code: The Most Beautiful False Theory in Biochemistry – Part 3
Does Nature use only 20 amino acids?
See similar articles by Klaus Roth published on ChemistryViews.org