Venomous animals such as snakes, spiders, or scorpions developed clever poison systems over the course of their evolution. These often contain a vast number of highly active biomolecules and are used to capture prey, as a defense, or to avoid competition. These venomous animals induce anxiety in many people since many of them can poison us with their toxins. At the same time, these creatures represent a treasure trove of highly promising biomolecules that could be useful in medicine.
In the first part of this article, we looked at venomous animals and their toxins. These can be used to fight disease. In the second part, we look at how animal toxins could also be used as biopesticides to help inhibit the spread of infectious diseases and at what the new field of “venomics” is all about.
3 Animal Toxins for the Control of Disease Carriers
In the wake of climate change and globalization, many animals are turning up as invasive species in areas outside of their natural habitat. A neophyte is a species that is not native to a geographical region and was just introduced in recent times. These often include dreaded carriers of infectious diseases, especially various species of mosquito (see Fig. 4). Together with their insect hosts, the diseases they carry, such as malaria, yellow fever, and dengue, are extending the range over which they spread, step by step [10].
|
|
|
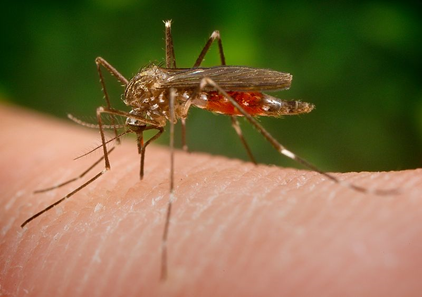
Figure 4. The Asian bush mosquito is a species native to Japan, Korea, and South China, that has been introduced as an invasive species in North America and Europe. It is a significant transmitter of pathogens like West Nile Virus and various types of encephalitis viruses in humans. |
 Traditional synthetic insecticides (see “Pyrethrum: History of a Bio-Insecticide“, article by Klaus Roth) were introduced to control harmful or dangerous insects. Unfortunately, these lack specificity, so their use can lead to a general reduction in the local insect population, including beneficial or protected species.
Traditional synthetic insecticides (see “Pyrethrum: History of a Bio-Insecticide“, article by Klaus Roth) were introduced to control harmful or dangerous insects. Unfortunately, these lack specificity, so their use can lead to a general reduction in the local insect population, including beneficial or protected species.
Furthermore, this is leading to an increase in resistance to synthetic insecticides. The control of disease carriers thus presents a major challenge for the future. Animal toxins may be a possible response to this challenge [7].
Insects are the primary source of food for venomous animals such as scorpions, assassin bugs, and particularly spiders. Their toxins have, thus, been optimized over millions of years to target specific insects. This makes their toxins outstanding starting materials for the development of new biopesticides that could replace old, tried-and-true synthetic formulations in the future.
One big advantage of animal venoms is the powerful specificity of some toxins. For example, toxins were isolated from the venoms of the desert bush spider (Digueta canities) and the African Golden Baboon (Augacephalus ezendami) that specifically work against the German cockroach (Blatella germanica) but do not affect other species of cockroach [11]. Another toxin, produced by the fusion of venom from the Australian funnel-web spider (Hadronyche versuta) with proteins from the snowdrop flower (Galanthus nivalis), is highly effective against a wide range of insect pests but not against honeybees [12].
Another big advantage of animal toxins over synthetic insecticides is their chemical properties. They are made of polypeptides, so they degrade over time in the environment and do not accumulate. Highly specific toxins with insecticidal effects represent a promising and environmentally friendly alternative to the substances available for the chemical control of insects that carry disease (“vector insects”), and thus, the fight against newly emerging infectious diseases [13].
4 A New Focus on Neglected Venomous Fauna
Research into animal venoms was long focused almost exclusively on specific groups of animals. Thus, the toxins of venomous snakes, followed by cone snails, scorpions, and the large tropical spider species were studied almost exclusively [14]. This was due on the one hand to an anthropocentric perspective on venoms: The basic assumption was often that the species dangerous to humans may produce particularly effective molecules in their venom. On the other hand, a profound understanding of these more dangerous toxins was particularly important for optimizing treatments for poisonings and producing antivenoms.
Another factor that kept the focus on these particular animal groups was the fact that it was possible to obtain the quantities of toxin required for biochemical analysis from them. The analytical methods of the past were very sample intensive and often required several milligrams of test substance [15]. These relatively large amounts could easily be obtained from venomous snakes by “milking”. However, most venomous animals are small and yield only tiny amounts of toxin. For this reason, only the venoms of large or dangerous species were studied intensively in the past, while relatively little is known about the toxins of most small and harmless European species.
5 The Young Research Field of “Venomics”
Today, the newest techniques for sequencing nucleic acids coupled with ultramodern mass spectrometers and supported by bioinformatics allow for the characterization of toxins from small species in a new integrated research field known as “venomics” [16]. The term venomics is derived from the English word for actively toxic animals (“venomous”) and the frequently used abbreviation for the high-throughput methods of systems biology, the “omics” technologies.
5.1 Analytical Methods
Whereas traditional methods used in animal toxin research started with large amounts of material and were required to individually analyze every component in the toxin being investigated, venomics experiments make it possible to identify all the components that can be measured by a given method at the same time. Most often, the proteins contained in the toxin are examined via proteomics (the analysis of all polypeptides present).
Another important method is transcriptomics (the analysis of all mRNAs present). This frequently involves sequencing and studying a tissue-specific transcriptome from the venom gland, either to derive the venom’s composition or to use the sequences obtained as a databank for subsequent proteomics experiments.
With increasing frequency, analyses of complete genomes are being used in animal toxin research, particularly in investigations of the evolution of toxins and gene regulation. In the future, this type of genome data could also serve as a basis for identifying new toxins with potential applications. It is, thus, very gratifying that the genomes of more and more toxic animals are becoming available. The latest examples include the Indian or spectacled cobra (Naja naja), the Komodo dragon (Varus komodoensisi), and the indigenous wasp spider (Argiope bruennischi) [17–19].
Another great advantage of venomics is that the toxins present in a given poison cocktail no longer need to be chemically purified. Because venomics experiments reveal the amino acid sequences of the toxins, biotechnological techniques can be used instead to produce larger amounts of the toxins in the laboratory. A wide range of methods is now available to achieve this. The most frequently used is heterologous expression in bacterial cells. However, in the recent past, technologies using eukaryotic cells or even fully cell-free methods were used to produce animal toxin components in the laboratory [20–22].
5.2 Major Progress
Although venomics is still a very young discipline, great progress has already been made. While next to nothing was known about smaller toxic animals until recently, our knowledge about the toxins from robber flies, assassin bugs, wasps, worms, and small spiders like the wasp spider is growing steadily, which makes it possible to obtain many valuable bioresources from the toxic animals that live in our backyards [23–28]. Some especially exciting examples of this are recently published papers about ant toxins.
Ant Venoms
Ant venoms evolved to protect the millions of inhabitants of a colony against scavengers and predators, especially mammals. One characteristic feature of ant venoms is that they cause extremely painful reactions. The Australian red bull ant (Myrmecia gullosa) is particularly notorious and can cause extreme pain that lasts for days. Australian researchers have recently been able to identify the toxin responsible. It is a molecular emulator of the epidermal growth factor (EGF) that is widespread among animals. It is a protein that stimulates cell growth and cell differentiation through binding to its receptor, EGFR. The EGF emulator from the venom binds to the natural receptors for this hormone and lingers there for an extremely long time [29].
It was previously not known that this receptor and its ligands play a role in mammalian pain. For pharmaceutical applications, this type of long-lasting interaction is very interesting, because we could use it to learn how to optimize the duration of effect for future drugs.
The application of venomics has also recently made it possible to examine the toxins of much smaller and significantly less painful stinging ant species from central Europe. Intriguingly, recent studies of two ant species, Myrmica rubra and Myrmica ruginodis, have shown that these distant relatives of the red bull ant produce comparable EGF-like toxins [30]. Understanding what effects these have and to what extent their binding strength deviates from that of the toxins in their Australian relatives could provide a key to understanding the impressive pharmacological activity of EGF-emulator toxins.
In addition, the venoms of both Myrmica species contain even more exciting components. For example, several toxins with structural features of antimicrobial peptides (AMPs) were found [31]. Very recently functional studies of these AMP-like toxins clearly demonstrated that they demonstrate broad-spectrum antibiotic effects against various bacteria, including several pathogens. One component in particular (U-MYRTX-Mrug5a) demonstrated truly potent activity against Listeria monocytogenes, the cause of listeriosis, an infectious disease that is triggered by the consumption of animal products and can be life-threatening for people with weakened immune systems.
Pseudoscorpion Venoms
Toxins from insects are not the only sources of new weapons in the fight against infectious diseases: Arachnids are also a highly promising source. Pseudoscorpions are among the lesser-known arachnids, not least because of their tiny size (only a few millimeters). These animals also have active toxin systems in appendages known as pedipalps. With venomics, it was recently possible to decode the toxins of several pseudoscorpions [32]. This led to the discovery of a class of toxins called checacins, which are also similar to AMPs.
Several examples from this class have now been tested, and checacin-1 appears to be especially intriguing for biomedicine. This toxin provokes a strong antibiotic effect, especially against methicillin-resistant Staphylococcus aureus—a major multiresistant hospital pathogen. At the same time, checacin-1 is almost not cytotoxic at all, making it a highly promising candidate for the development of new antibiotics [33].
Further Reading
The investigation of venoms and their potential applications is an enormous undertaking; we recommend further literature on the subject for interested readers [34,35].
6 Take-Home Messages
Venomous animals inject their toxins, whereas passively toxic animals transmit their poison via membrane transport.
Animal toxins are widespread in the animal kingdom and have been evolutionarily optimized over millions of years with regard to their function.
Animal toxins are among the most complex chemical systems found in the animal kingdom. Excellent use can be made of the bioactivity, specificity, and selectivity of animal toxins in deriving innovative drugs from them in the future.
In the past, primarily dangerous, tropical toxic animals were studied; harmless, native species have only recently entered the spotlight with the development of improved analytical methods.
References
[10] J. Schmidt-Chanasit, J. Tiedke, Tropische Viren im Anflug nach Europa, Med. Monatsschr. Pharm. 2017, 40, 151–153.
[11] N. S. Bende, S. Dziemborowicz, M. Mobli, V. Herzig, J. Gilchrist, J. Wagner, G. M. Nicholson, G. F. King, F. Bosmans, A distinct sodium channel voltage-sensor locus determines insect selectivity of the spider toxin Dc1a, Nat. Commun. 2014, 5, 4350. https://doi.org/10.1038/ncomms5350
[12] E. Y. T. Nakasu, S. M. Williamson, M. G. Edwards, E. C. Fitches, J. A. Gatehouse, G. A. Wright, A. M. R. Gatehouse, Novel biopesticide based on a spider venom peptide shows no adverse effects on honeybees, Proc. Royal Soc. B 2014, 281, 20140619. https://doi.org/10.1098/rspb.2014.0619
[13] V. Herzig, B. Cristofori-Armstrong, M. R. Israel, S. A. Nixon, I. Vetter, G. F. King, Animal toxins — Nature’s evolutionary-refined toolkit for basic research and drug discovery, Biochem. Pharmacol. 2020, 181, 114096. https://doi.org/10.1016/j.bcp.2020.114096
[14] B.Marcus Von Reumont, L. I. Campbell, R. A. Jenner, Quo Vadis Venomics? A Roadmap to Neglected Venomous Invertebrates, Toxins 2014, 6, 3488–3551. https://doi.org/10.3390/toxins6123488
[15] T. Lüddecke, A. Vilcinskas, S. Lemke, Phylogeny-Guided Selection of Priority Groups for Venom Bioprospecting: Harvesting Toxin Sequences in Tarantulas as a Case Study, Toxins 2019, 11, 488. https://doi.org/10.3390/toxins11090488
[16] S. H. Drukewitz, B. M. von Reumont, The Significance of Comparative Genomics in Modern Evolutionary Venomics, Frontiers Ecol. Evol. 2019, 7, 163. https://doi.org/10.3389/fevo.2019.00163
[17] K. Suryamohan, S. P. Krishnankutty, J. Guillory, M. Jevit, M. S. Schröder, M. Wu, B. Kuriakose, O. K. Mathew, R. C. Perumal, I. Koludarov, L. D. Goldstein, K. Senger, M. Davis Dixon, D. Velayutham, D. Vargas, S. Chaudhuri, M. Muraleedharan, R. Goel, Y.-J. J. Chen, A. Ratan, P. Liu, B. Faherty, G. de la Rosa, H. Shibata, S. Seshagiri, The Indian cobra reference genome and transcriptome enables comprehensive identification of venom toxins, Nat. Genet. 2020, 52, 106–117. https://doi.org/10.1038/s41588-019-0559-8
[18] A. L. Lind, Y. Y. Y. Lai, Y. Mostovoy, A. K. Holloway, A. Iannucci, A. C. Y. Mak, M. Fondi, V. Orlandini, W. L. Eckalbar, M. Milan, M. Rovatsos, I. G. Kichigin, A. I. Makunin, M. Johnson Pokorná, M. Altmanová, V. A. Trifonov, E. Schijlen, L. Kratochvíl, R. Fani, P. Velenský, I. Rehák, T. Patarnello, T. S. Jessop, J. W. Hicks, O. A. Ryder, J. R. Mendelson III, C. Ciofi, P.-Y. Kwok, K. S. Pollard, B. G. Bruneau, Genome of the Komodo dragon reveals adaptations in the cardiovascular and chemosensory systems of monitor lizards, Nat. Ecol. Evol. 2019, 3, 1241–1252. https://doi.org/10.1038/s41559-019-0945-8
[19] M. M. Sheffer, A. Hoppe, H. Krehenwinkel, G. Uhl, A. W. Kuss, L. Jensen, C. Jensen, R. G. Gillespie, K. J. Hoff, S. Prost, Chromosome-level reference genome of the European wasp spider Argiope bruennichi: a resource for studies on range expansion and evolutionary adaptation, Gigascience 2021, 10, giaa148. https://doi.org/10.1093/gigascience/giaa148
[20] J. K. Klint, S. Senff, N. J. Saez, R. Seshadri, H. Y. Lau, N. S. Bende, E. A. B. Undheim, L. D. Rash, M. Mobli, G. F. King, Production of Recombinant Disulfide-Rich Venom Peptides for Structural and Functional Analysis via Expression in the Periplasm of E. coli, PloS ONE 2013, 8, e63865. https://doi.org/10.1371/journal.pone.0063865
[21] H. C. da Justa, F. Hitomi Matsubara, E. de-Bona, Z. Schemczssen-Graeff, N. L. Costacurta Polli, T. L. de Mari, M. Boia-Ferreira, J. C.Minozzo, A. C. Martins Wille, A. Senff-Ribeiro, L. H. Gremski, S. Sanches Veiga, LALLT (Loxosceles Allergen-Like Toxin) from the venom of Loxosceles intermedia: Recombinant expression in insect cells and characterization as a molecule with allergenic properties, Int. J. Biol. Macromol. 2020, 164, 3984–3999. https://doi.org/10.1016/j.ijbiomac.2020.08.212
[22] T. Lüddecke, A. Paas, L. Talmann, K. N. Kirchhoff, B. M. von Reumont, A. Billion, T. Timm, G. Lochnit, A. Vilcinskas, A Spider Toxin Exemplifies the Promises and Pitfalls of Cell-Free Protein Production for Venom Biodiscovery, Toxins 2021, 13, 575. https://doi.org/10.3390/toxins13080575
[23] S. H. Drukewitz, L. Bokelmann, E. A. B. Undheim, B. M. von Reumont, Toxins from scratch? Diverse, multimodal gene origins in the predatory robber fly Dasypogon diadema indicate a dynamic venom evolution in dipteran insects, GigaScience 2019, 8, giz081. https://doi.org/10.1093/gigascience/giz081
[24] M. Tonk, A. Vilcinskas, C. G. Grevelding, S. Haeberlein, Anthelminthic Activity of Assassin Bug Venom against the Blood Fluke Schistosoma mansoni, Antibiotics 2020, 9, 664. https://doi.org/10.3390/antibiotics9100664
[25] R. Özbek, N. Wielsch, H. Vogel, G. Lochnit, F. Foerster, A. Vilcinskas, B. M. von Reumont, Proteo-Transcriptomic Characterization of the Venom from the Endoparasitoid Wasp Pimpla turionellae with Aspects on Its Biology and Evolution, Toxins 2019, 11, 721. https://doi.org/10.3390/toxins11120721
[26] B. M. von Reumont, T. Lüddecke, T. Timm, G. Lochnit, A. Vilcinskas, J. von Döhren, M. A. Nilsson, Proteo-Transcriptomic Analysis Identifies Potential Novel Toxins Secreted by the Predatory, Prey-Piercing Ribbon Worm Amphiporus lactifloreus, Marine Drugs 2020, 18, 407. https://doi.org/10.3390/md18080407
[27] T. Lüddecke, B. M. von Reumont, F. Förster, A. Billion, T. Timm, G. Lochnit, A. Vilcinskas, S. Lemke, An Economic Dilemma between Molecular Weapon Systems May Explain an Arachno-Atypical Venom in Wasp Spiders (Argiope bruennichi), Biomolecules 2020, 10, 978. https://doi.org/10.3390/biom10070978
[28] B. M von Reumont, G. Anderluh, A. Antunes, N. Ayvazyan, D. Beis, F. Caliskan, A. Crnković, M. Damm, S. Dutertre, L. Ellgaard, G. Gajski, H. German, B. Halassy, B.-F. Hempel, T. Hucho, N. Igci, M. P. Ikonomopoulou, I. Karbat, M. I. Klapa, I. Koludarov, J. Kool, T. Lüddecke, R. Ben Mansour, M. V. Modica, Y. Moran, A. Nalbantsoy, M. E. Pachón Ibáñez, A. Panagiotopoulos, E. Reuveny, J. Sánchez Céspedes, A. Sombke, J. M. Surm, E. A. B. Undheim, A. Verdes, G. Zancolli, Modern venomics—Current insights, novel methods, and future perspectives in biological and applied animal venom research, GigaScience 2022, 11, giac048. https://doi.org/10.1093/gigascience/giac048
[29] D. A. Eagles, N. J. Saez, B. Krishnarjuna, J. J. Bradford, Y. K.-Y. Chin, H. Starobova, A. Mueller, M. E. Reichelt, E. A. B. Undheim, R. S. Norton, W. G. Thomas, I. Vetter, G. F. King, S. D. Robinson, A peptide toxin in ant venom mimics vertebrate EGF-like hormones to cause long-lasting hypersensitivity in mammals, Proc. Natl. Acad. Sci. USA 2022, 119, e2112630119. https://doi.org/10.1073/pnas.2112630119
[30] S. Hurka, K. Brinkrolf, R. Özbek, F. Förster, A. Billion, J. Heep, T. Timm, G. Lochnit, A.Vilcinskas, T. Lüddecke, Venomics of the Central European Myrmicine Ants Myrmica rubra and Myrmica ruginodis, Toxins 2022, 14, 358. https://doi.org/10.3390/toxins14050358
[31] S. Hurka, T. Lüddecke, A. Paas, L. Dersch, L. Schulte, J. Eichberg, K. Hardes, K. Brinkrolf, A. Vilcinskas, Bioactivity Profiling of In Silico Predicted Linear Toxins from the Ants Myrmica rubra and Myrmica ruginodis, Toxins 2022, 14, 846. https://doi.org/10.3390/toxins14120846
[32] J. Krämer, H. Pohl, R. Predel, Venom collection and analysis in the pseudoscorpion Chelifer cancroides (Pseudoscorpiones: Cheliferidae), Toxicon 2019, 162, 15–23. https://doi.org/10.1016/j.toxicon.2019.02.009
[33] J. Krämer, T. Lüddecke, M. Marner, E. Maiworm, J. Eichberg, K. Hardes, T. F. Schäberle, A. Vilcinskas, R. Predel, Antimicrobial, Insecticidal and Cytotoxic Activity of Linear Venom Peptides from the Pseudoscorpion Chelifer cancroides, Toxins 2022, 14, 58. https://doi.org/10.3390/toxins14010058
[34] G. Habermehl, Gift-Tiere und ihre Waffen, Springer-Verlag, Berlin-Heidelberg, Germany, 1994. ISBN: 978-3-540-56897-1
[35] D. Mebs, Gifttiere: Ein Handbuch für Biologen, Toxikologen, Ärzte und Apotheker, Wissenschaftliche Verlagsgesellschaft Stuttgart, Germany, 1992. ISBN: 978-3-8047-2510-2
The Authors
Tim Lüddecke, born in 1989, trained as a chemical laboratory assistant from 2008 to 2012. From 2012 until 2018, he studied biology at TU Braunschweig, Germany, with a concentration in biochemistry. From 2018 to 2021, he completed his doctorate at the Fraunhofer Institute for Molecular Biology and Applied Ecology IME in Gießen, Germany. Since 2018 he has additionally been an Associate Scientist at the LOEWE Centre for Translational Biodiversity Genomics in Frankfurt am Main, Germany. Since 2019 he has been co-chairman of the German Society of Arachnology and from 2020 to 2021, he was a German representative in the Management Committee (substitute) of the EU COST Action “European Venom Network”. Since 2021 he has been a Group Leader in the Division of Animal Venomics of the Fraunhofer Institute for Bioresources, currently under construction in Gießen.
Andreas Vilcinskas, born in 1964, studied biology at TU Kaiserslautern and the Free University of Berlin, Germany. He received his doctorate in 1994 from the Institute for Zoology at the Free University of Berlin and became a Professor there in 1998 in the field of zoology. From 1994 until 2004 he was a Visiting Professor and Acting Chair of Evolutionary Biology and Special Zoology at the Institute for Biochemistry and Biology at the University of Potsdam. In 2004 he was appointed Professor of Applied Entomology at the Justus Liebig University in Gießen and founded the world’s first institute for insect biotechnology. He is currently establishing the Fraunhofer Institute for Bioresources in Gießen, where animal toxins will be developed as resources for new pharmaceuticals.
Modified, extended translation of:
- Heilen mit Tiergiften,
Tim Lüddecke, Andreas Vilcinskas,
Chem. Unserer Zeit 2021.
https://doi.org/10.1002/ciuz.202100005
Translated by Caroll Pohl-Ferry
Healing with Venom – Part 1
Venomous animals and their poisons
See similar articles by Klaus Roth published in ChemistryViews




I have always thought that the answer to every ailment is already sitting in nature. The science of venomics is hinted at by Linus Pauling’s notion of orthomolecular medicine. It is the idea that most disease states can be cured or at least mitigated by the appropriate supplementation.
This is an exciting field because the high affinity and superb specificity of venom, poisons and oligopeptide toxins could steer us in the direction of many discoveries in the area of structure activity correlations. It is the next iteration necessary for the waning efficacy of antibiotics.